(C) 2012 Milton Groppo. This is an open access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC-BY), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
For reference, use of the paginated PDF or printed version of this article is recommended.
The position of the plant genus Pitavia within an infrafamilial phylogeny of Rutaceae (rue, or orange family) was investigated with the use of two non-coding regions from cpDNA, the trnL-trnF region and the rps16 intron. The only species of the genus, Pitavia punctata Molina, is restricted to the temperate forests of the Coastal Cordillera of Central-Southern Chile and threatened by loss of habitat. The genus traditionally has been treated as part of tribe Zanthoxyleae (subfamily Rutoideae) where it constitutes the monogeneric tribe Pitaviinae. This tribe and genus are characterized by fruits of 1 to 4 fleshy drupelets, unlike the dehiscent fruits typical of the subfamily. Fifty-five taxa of Rutaceae, representing 53 genera (nearly one-third of those in the family) and all subfamilies, tribes, and almost all subtribes of the family were included. Parsimony and Bayesian inference were used to infer the phylogeny; six taxa of Meliaceae, Sapindaceae, and Simaroubaceae, all members of Sapindales, were also used as out-groups. Results from both analyses were congruent and showed Pitavia as sister to Flindersia and Lunasia, both genera with species scattered through Australia, Philippines, Moluccas, New Guinea and the Malayan region, and phylogenetically far from other Neotropical Rutaceae, such as the Galipeinae (Galipeeae, Rutoideae) and Pteleinae (Toddalieae, former Toddalioideae). Additionally, a new circumscription of the subfamilies of Rutaceae is presented and discussed. Only two subfamilies (both monophyletic) are recognized: Cneoroideae (including Dictyolomatoideae, Spathelioideae, Cneoraceae, and Ptaeroxylaceae) and Rutoideae (including not only traditional Rutoideae but also Aurantioideae, Flindersioideae, and Toddalioideae). As a consequence, Aurantioideae (Citrus and allies) is reduced to tribal rank as Aurantieae.
Biogeography, Cneoroideae, phylogeny, Pitavia, Rutaceae, Rutoideae, rps16, subfamily, trnL-trnF
Rutaceae is a large, predominantly tropical and subtropical family, consisting of 150–162 genera and 1500–2096 species, with three main centers of diversity: Tropical America, southern Africa, and Australia (
Given its great morphological diversity that include a variety of habits, flowers, and fruits, allied with a broad geographic distribution, Rutaceae has been traditionally divided into subfamilies, tribes, and subtribes, following the classifications of
One subtribe that had not yet been sampled in molecular phylogenetic studies of the family is Pitaviinae, which comprises a single genus and species, Pitavia punctata Molina. (photos can be seen at http://www.florachilena.cl/Niv_tax/Angiospermas/Ordenes/Sapindales/Rutaceae/Pitavia%20punctata/Pitavia%20punctata.htm ). This species is restricted to the temperate forests of the Coastal Cordillera of south-central Chile at 35°-38°S. It is the sole species of Rutaceae native to the continental area of that country (another rutaceous species, Zanthoxylum mayu Bertero, is restricted to Juan Fernández Island, see
Pitavia consists of small trees with simple, opposite to whorled leaves and unisexual, 4-merous, diplostemonous flowers of which the four carpels are proximally connate and joined subapically in a common style. The fruit is composed of one to four fleshy drupes, each with a solitary seed (
The main objective of this study is to determine the position of Chilean Pitavia within a the Rutaceae phylogeny and to assess whether the genus is more closely related to Australasian members of Rutaceae or to Neotropical ones (such as the tribes Galipeeae or Toddalioideae, in Engler's [1931] classification). To examine this, we chose two non-coding regions from cpDNA, the rps16 gene intronand the trnL-trnF region, for a representative sampling of Rutaceae. The type II rps16 intron was first used for phylogenetic analysis by
The present study includes genera from all Englerian subfamilies, tribes, and almost all subtribes of Rutaceae and accounts for a broad geographic representation of the family. The large sample of genera used here provides a basis for revising Engler’s circumscription of the subfamilies of Rutaceae. Although new arrangements of subfamilies have been proposed (e.g.,
Given the uncertain position of Pitavia within the Rutaceae phylogeny, representatives of all subfamilies and tribes and almost all subtribes of Rutaceae (sensu
Total genomic DNA was extracted from 5 mg of dried leaf sample from a collection of Pitavia punctata (Kubitzki 01-07, Herbarium HBG) using a modified CTAB protocol (
Automated alignments of the sequences were made with Clustal X (
Bayesian phylogenetic inference was performed with MrBayes v. 3.1.2 (
The Akaike Information Criterion implemented in MrModelTest chose the GTR + G + I evolutionary model as the best fit for the rps16 and trnL-trnF concatenated sequences. The burn-in value was set to 4, 000 tree samplings, reflecting 2 million generations, i.e., long after the analysis was considered to have stabilized (by inspection of effective sample sizes and standard deviation of split frequencies). The aligned matrix comprised a total of 2, 229 characters: 1123 invariable, 467 variable but parsimony-uninformative, and 639 parsimony-informative. At the point when the search was interrupted, parsimony analysis resulted in 10, 000 most parsimonious trees with 2, 535 steps, consistency index (CI) = 0.68 (0.51 excluding uninformative characters), and retention index (RI) = 0.68.
The majority-rule consensus tree with posterior probabilities (PP) that was estimated using Bayesian Inference is shown in Fig. 1. Bootstrap percentages (BP) are also shown for clades recovered in the majority-rule consensus tree of the bootstrap analysis. As commonly seen in the literature, a higher resolution was obtained with the Bayesian analysis than with the majority-rule bootstrap consensus trees based on parsimony, as can be noted in the figure: many clades that appeared in the 50% majority-rule Bayesian tree do were not recovered in the boostrap analysis. (even in clades with PP as high as 0.99). Given its better resolution and branch support values, and the generally accepted superiority of Bayesian methods in inferring reliable phylogenetic relationships we chose to discuss our results on the basis of the Bayesian tree.
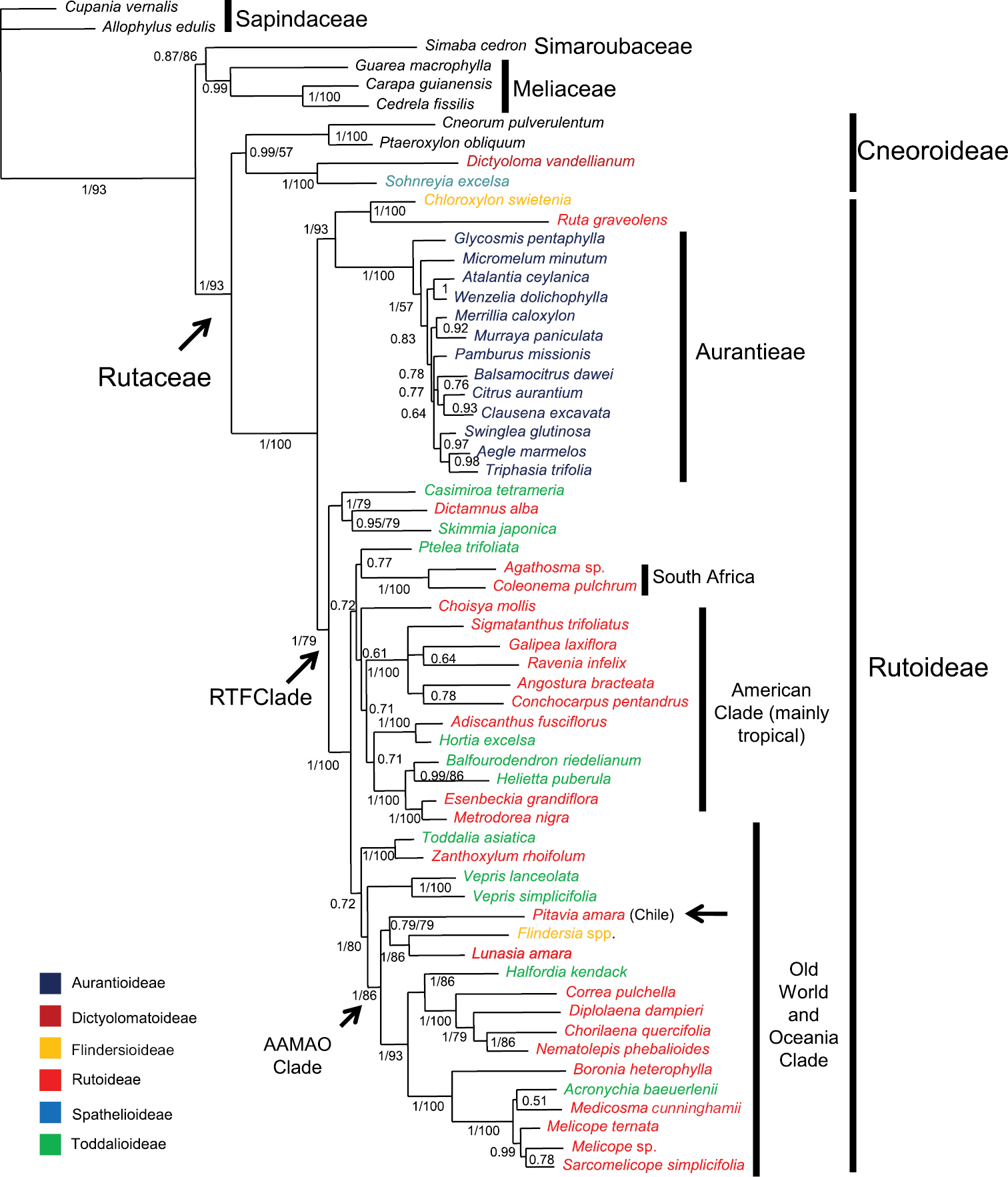
Majority-rule consensus tree of Rutaceae estimated using Bayesian inference on a combined rps16 and trnL-trnF dataset. Posterior probabilities (PP ≥50%) are shown above branches. Bootstrap percentages (BP, only for branches in agreement with those obtained in the Bayesian analysis) follow posterior probabilities; when only one number appears supporting a clade it refers to Bayesian posterior probabilities. Taxon names are color-coded to indicate their Englerian assignment to subfamilies. A new proposal that recognizes monophyletic groups (subfamilies Cneoroideae and Rutoideae and tribe Aurantieae) is indicated by the vertical bars. The position of Pitavia punctata, as well as the Rutaceae, the “RTF” (from Rutoideae, Toddalioideae and Flindersia) and “AAMAO” (”African-Asian-Malesian-Astralasian-Oceanic”) clades (see text) are indicated by arrows. Note: Zanthoxylum is pantropical.
Topology of the Bayesian analysis was congruent with that obtained in the study of Groppo et al.(2008) using parsimony, but with a better resolution of some clades as discussed above. Rutaceae appeared as monophyletic (PP=1, BP=93), encompassing two internal clades, one with Cneorum, Ptaeroxylon, Sohnreyia, and Dictyoloma with mixed support (strong PP=0.99 and weak BP=57) and another with the remaining Rutaceae (1/100). This clade is divided in two sister-groups: one (1/93) formed by Chloroxylon (Flindersioideae) plus Ruta (Rutoideae) and all Aurantioideae (the only monophyletic Englerian subfamily with more than one genus) and the other (1/79) with interdigitated representatives of Rutoideae (without Ruteae), Toddalioideae, and Flindersia (Flindersioideae), the RTF Clade. Chilean Pitavia appears as part of this last group, in a clade containing also Lunasia and Flindersia (1/86), which in turn is part of a larger clade (1/86) formed by representatives of Rutaceae of Old World, Australasia and Oceanic islands from Pacific (the ”African-Asian-Malesian-Astralasian-Oceanic - AAMAO clade”)
A clade comprising Flindersia and Lunasia was obtained also by
The morphological resemblance of Pitavia to Rutaceae from Oceania and Southern Asia was implicitly suggested by
The association of Chilean Pitavia with Flindersia and Lunasia is an example of biogeographical affinity between components of the faunas and floras occurring on both sides of the Pacific Ocean especially in the Southern Hemisphere (for a reviews of this issue and examples, see
Distributional patterns in Rutaceae have often been explained on basis of vicariance events (e.g.,
Estimates of the age of Rutaceae based on molecular studies vary from 37 to 93.3 mybp (
Despite the linking of Flindersia, Lunasia, and Pitavia shown in the present study, it is premature to say that Pitavia is indeed sister to the Flindersia/Lunasia clade because many genera, such as Dinosperma, Perryodendron, Pitaviaster, Crossosperma, and Dutailliopsis, have not yet been included in phylogenetic studies. However, the support (PP PP, 86% of BP) of the clade (Pitavia (Flindersia, Lunasia)) is strong enough to refute an association of Pitavia with the clade with Acronychia, Melicope, and Sarcomelicope (as suggested by
The placement of Chloroxylon (Flindersioideae) near Ruta is supported by the possession of diplostemonous flowers, unguiculate petals with concavities embracing the smaller antepetalous stamens, a developed urceolate disc, and more than two ovules per locule. The base chromosome number in both genera is X=10 (
Another objective of this work is to present a new proposal to replace the standard classification of subfamilies of Rutaceae, replacing that proposed by
Given these data, realignments of the infrafamilial groups in Rutaceae have been recently made. In a survey of secondary metabolites (largely influenced by the conclusions of
The remaining Rutaceae or “core Rutaceae” (
A summary of the new circumscription of subfamilies in Rutaceae proposed in this study. Approximate number of genera and species and distribution taken from
| Subfamily | Circumscription | Approximate number of genera/ (species) | Distribution |
|---|---|---|---|
| Cneoroideae | Englerian Dictyolomatoideae and Spathelioideae ( |
8 (35) | Pantropical, extending to subtropical regions in Europe |
| Rutoideae | Englerian Aurantioideae, Flindersioideae, Rutoideae, and Toddalioideae ( |
114 (1770) in clade RTF; 26 (206) in Aurantieae; 7 (84) in Ruteae (including Cneoridium and Haplophyllum), plus 1 (1) in Chloroxylon. Total: 148 (2061) | Pan-tropical, some in temperate or desert areas worldwide |
| Total | 156 (2096) |
Restricting the name Rutoideae to Ruta and its allies in tribe Ruteae (excluding Dictamnus, see
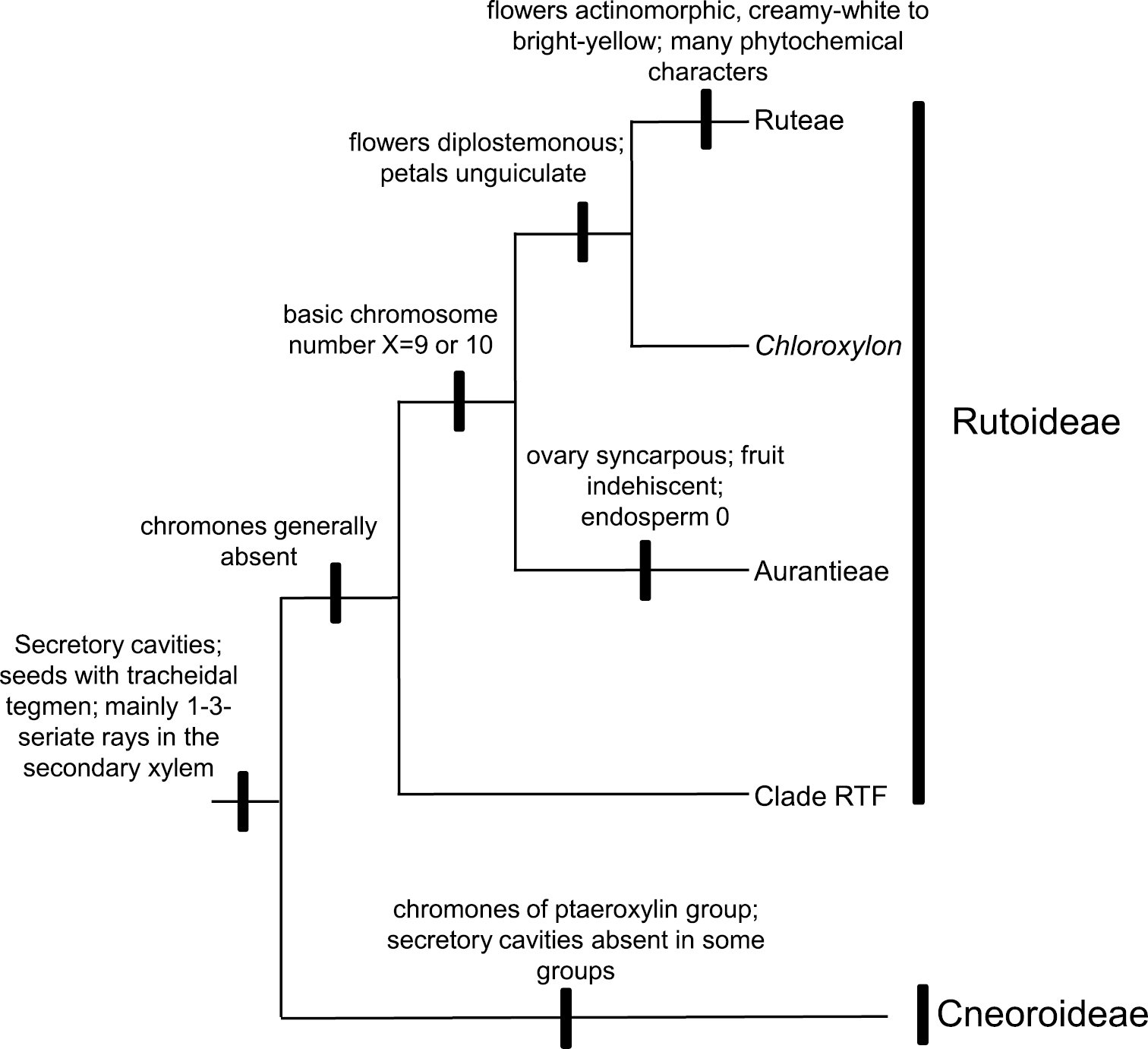
A summary of the phylogenetic relationships of proposed subfamilies in Rutaceae, with some non-molecular characteristics plotted onto a simplified cladogram based on the Bayesian tree. Chloroxylon is doubtfully attached to Aurantieae. Haplophyllum and Cneoridium (both from Englerian Ruteae but closer to Aurantieae that to remaining Ruteae, see
Formal recognition of the expanded Rutoideae and the Cneoroideae at the family level (i.e., respectively as Rutaceae sensu stricto and Cneoraceae) is at odds with shared morphological characters. One synapomorphy uniting these two clades, despite its absence in some Cneoroideae (due to a secondary loss,
The classification scheme presented here, with only two monophyletic subfamilies, Cneoroideae and Rutoideae, is a framework for further studies of the family. The next step is the re-circumscription of groups below the subfamilial level, i.e., the tribes and subtribes, as
The authors thank Thomas G. Hartley for sending samples of Australian plants (Acronychia, Flindersia, Halfordia, Melicope, Sarcomelicope) to MG; the Margareth Mee Foundation for a grant to MG that allowed him to visit European herbaria and the Jodrell Laboratory at Kew; Edith Kapinos, responsible for the plant DNA bank at the Jodrell Laboratory, for her help and technical support; Klaus Kubitzki for sending a sample of Pitavia and for a critical reading of an earlier version of the manuscript, as well as two anonymous reviewers for revising the text; Tatiana Parreiras Martins for sequencing procedures. This work was supported by grants to MG from FAPESP (Fundação de Amparo à Pesquisa do Estado de São Paulo; grants #2000/07401-0, 2006/03170-0, 2009/54031-0, 2011/19446-0) and from CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico). AA is supported by grants from the Swedish and the European Research Councils.
Summary of the subfamilies recognized in the present circumscription of Rutaceae. Source of dates of publication is Reveal, J. “Suprafamilial Names of Extant Vascular Plants, ” available at http://www.plantsystematics.org/reveal/pbio/fam/hightaxa7.html
(for a complete list of synonyms see
Rutaceae A.Juss., Gen Pl.: 296. 4 Ago 1789. (type: Ruta L.) nomen conservandum
Subfamily Cneoroideae Webb, London J. Bot. 1: 257. 1 Mai 1842. (Cneoreae). (type: Cneorum L.) [Circumscription: Englerian Dictyolomatoideae and Spathelioideae (Engler 1931) and Cneoraceae and Pteroxylaceae (see
Synomyms:
Cneoraceae Vest, Anleit. Stud. Bot.: 267. 1818. (type: Cneorum L.)
Dictyolomatoideae (“Dictyolomoideae”) Engl. in Engl. and Prantl, Nat. Pflanzenfam. III, 4: 111. Mar 1896. (type: Dictyoloma A. Juss.).
Ptaeroxylaceae J.-F.Leroy, J. Agric. Trop. Bot. Appl. 7: 456. 1960. (type: Ptaeroxylon Radlk.).
Spathelioideae Engl. in Engl. and Prantl, Nat. Pflanzenfam. III, 4: 111. Mar 1896. (type: Spathelia L.).
Subfamily Rutoideae Arn., Encycl. Brit., ed. 7, 5: 104. 9 Mar 1832 (Ruteae). (type: Ruta L.) [Circumscription: Englerian Aurantioideae, Flindersioideae, Rutoideae, and Toddalioideae, see Engler (1931)]
Synomyms:
Aurantioideae Eaton, Bot. Dict., ed. 4: 39. Apr-Mai 1836 (Aurantiaceae). [(type: Aurantium L. (=Citrus L.)]. Recognized here as tribe Aurantieae Rchb., Fl. Germ, Excurs. 292). 840. 1832.
(note: see
Flindersiaceae C.T.White ex Airy Shaw, Kew Bull. 18: 257. 8 Dec 1964. (type: Flindersia R. Br.).
Flindersioideae Luerss., Handb. Syst. Bot. 2: 681. Jun 1881 (“Flindersieae”). (type: Flindersia R. Br.).
Toddalioideae K.Koch, Dendrologie 1: 564. 1869 (“Toddalieae”). (type: Toddalia A.Juss.).
Voucher information and Genbank accession numbers for taxa used in this study. Missing sequences are marked by “—.“ Arrangement of terminals in Rutaceae follows the new subfamilial delimitation proposed here. For classification of terminals in Engler (1931) and
Genbank accessions: trnL-trnF, rps16.
Rutaceae
Subfamily Cneoroideae
Cneorum pulverulentum Vent. — EU853787, EU853733. Dictyoloma vandellianum A. Juss. — EU853793, EU853739. Ptaeroxylon obliquum (Thunb.) Radlk. — EU853812, EU853762. Sohnreyia excelsa Krause — EU853820, EU853770.
Subfamily Rutoideae
Acronychia baeuerlenii T. G. Hartley — EU853774, EU853719. Adiscanthus fusciflorus Ducke — EU853775, EU853721. Aegle marmelos (L.) Corrêa ex Roxb. — AY295294, AY295268. Afraegle paniculata (Schum. and Thonn.) Engl. — AY295295, AY295269. Agathosma sp. — EU853776, EU853722. Angostura bracteata (Engl.) Kallunki — EU853778, EU853724. Atalantia ceylanica (Arn.) Oliv. — AY295288, AY295262. Balfourodendron riedelianum (Engl.) Engl. — EU853779, EU853725. Balsamocitrus dawei Stapf — AY295278, AY295252. Boronia heterophylla F. Muell. — EU853780, EU853726. Casimiroa tetrameria Millsp. — EU853782, EU853728. Chloroxylon swietenia DC. — AY295276, AY295250. Choisya mollis Standl. — EU853784, EU853730. Chorilaena quercifolia Endl. — EU853785, EU853731. Citrus aurantium L. — EU853786, EU853732. Clausena excavata Burm. f. — AY295284, AY295258. Coleonema pulchrum Hook. — EU853788, EU853734. Conchocarpus pentandrus (A.St.-Hil.) Kallunki and Pirani — EU853735, EU853735. Correa pulchella Mackay ex Sweet — EU853790, EU853736. Dictamnus albus L. — EU853792, EU853738. Diplolaena dampieri Desf. — EU853794, EU853740. Esenbeckia grandiflora Mart. — EU853795, EU853741. Flindersia australis R. Br. — AF038628 (intron) AF026009 (spacer), —, — spacer. Flindersia collina F. M. Bailey — —, EU853742. Galipea laxiflora Engl. — EU853796, EU853743. Halfordia kendack (Montrouz.) Guillaumin — EU853798, EU853745. Hortia excelsa Ducke — EU853801, EU853748. Glycosmis pentaphylla (Retz.) Corrêa — AY295279, AY295253. Helietta puberula R. E. Fries — EU853799, EU853746. Lunasia amara Blanco — EU853805, EU853753. Medicosma cunninghamii (Hook.) Hook. f. — EU853806, EU853754. Melicope ternata J. R. Forst. and G. Forst. — EU853808, EU853756. Melicope sp. — EU853807, EU853755. Metrodorea nigra A. St.-Hil. — EU853809, EU853757. Micromelum minutum (G. Forst.) Wight and Arn. — AF025520, AF320266. Murraya paniculata (L.) Jack — EU853810, EU853758. Nematolepis phebalioides Turcz. — AF025522 (spacer), EU853759. Pitavia amara Blanco —KC261635, KC261636. Ravenia infelix Vell. — EU853814, EU853764. Ruta graveolens L. — EU853815, EU853765. Pamburus missionis (Wight) Swingle — AY295300, AY295274. Ptelea trifoliata L. — EU853813, EU853763. Sarcomelicope simplicifolia (Endl.) T. G. Hartley — EU853816, EU853766. Sigmatanthus trifoliatus Huber ex Emmerich — EU853817, EU853767. Skimmia japonica Thunb. — EU853819, EU853769. Swinglea glutinosa (Blanco) Merr. — AY295285, AY295259. Toddalia asiatica (L.) Lam. — EU853821, AF320278. Triphasia trifolia (Burm. f.) P. Wilson — EU853822, AY295271. Vepris lanceolata (Lam.) G. Don — EU853823, EU853771. Vepris simplicifolia (Engl.) W. Mziray — EU853824, EU853772. Wenzelia dolichophylla (Lauterb. and K. Schum.) Tanaka — AY295286, AY295260. Zanthoxylum rhoifolium Lam. — EU853773, EU853720.
Meliaceae
Carapa guianensis Aubl. — EU853781, EU853727. Cedrela fissilis Vell. — EU853783, EU853729. Guarea macrophylla Vahl — EU853797, EU853744.
Sapindaceae
Allophylus edulis (A. St.-Hil.) Niederl. — EU853777, EU85853723. Cupania vernalis Cambess. — EU853791, EU853737.
Simaroubaceae